 
The Qscore assigned to each nucleotide tends to be low when a wrong nucleotide is assigned, as expected. S1), we plotted the distribution of correct/error nucleotides according to the Qscore returned by Oxford nanopore's basecaller, Guppy. 1A (and a normalized version in Supplementary Fig. In this data set, we can confidently attribute mismatches between the read sequences and the known wild-type as sequencing errors and matches as correct reads. We used a nanopore sequencer to read 5,847 strands of the wild-type KlenTaq gene (length 1,662 nucleotides), for which we have a ground-truth sequence obtained by Sanger sequencing ( Supplementary Sequence S1). This translates into a more efficient exploitation of the sequencing throughput. Therefore, as few as 5 to 7 reads return a trustable consensus sequence, outperforming the state-of-the-art tools for consensus computation of nanopore sequencing, Medaka and Nanopolish. 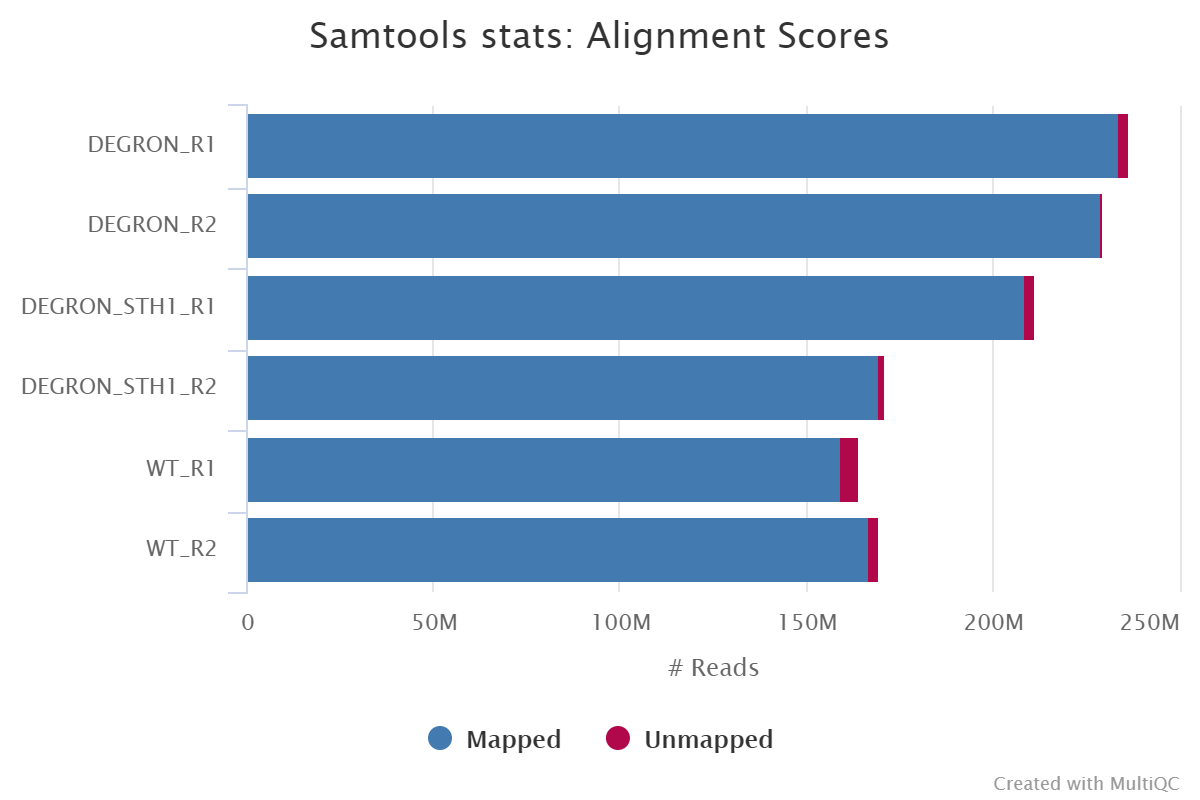
SINGLe reduced the sequencing noise, allowing a better identification of true point mutations. We first tested it on a small set of 7 known mutants containing 2 to 9 point mutations and later a larger library of approximately 1,200 variants. 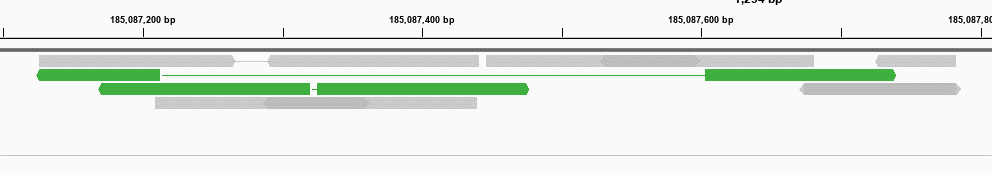
Here, we applied SINGLe to the gene of KlenTaq, a truncated variant of the well-known Taq polymerase, of approximately 1.7 kb in length. Finally, these values can be used in the consensus calling of individual variants. SINGLe is first trained on a set of reads of the reference by a nanopore sequencer, and then it is applied to the reads of the actual library to correct their quality scores (Qscore). In this work, we introduce SINGLe (SNPs In Nanopore reads of Gene Libraries), a method that improves detection of single mutations via consensus calling in reads of libraries for which a reference sequence is known. Inconveniently, these methods reduce the number of different variants that can be studied, because a part of the sequencing throughput is invested in reading each sequence multiple times. This has been achieved by creating sequence concatenates using rolling circular amplification, which retrieved an accuracy of 99.5% for coverage of 150×, and via gene barcoding prior to amplification with a reported accuracy over 99.9% for 25× coverage. These strategies aim to read and associate several replicates of the same molecule. While these tools primarily apply to genome assembly, a number of experimental protocols were developed in order to apply these pipelines in the specific case of amplicon library sequencing. Nanopolish reports an accuracy over 99.5%, for a 29× sequencing coverage, and Medaka 98% in detection of single-nucleotide polymorphisms (SNPs) with a coverage of 100×. These approaches start from a draft assembly and use the coverage depth to compute an averaged consensus at each position, via various computational approaches. Previous work aiming at high-quality sequencing from nanopore data has concentrated on polishing tools such as Nanopolish or Racon combined with Medaka. Unfortunately, nanopore's relatively high error rate (≈6–15%) prevents the accurate detection of point genetic variation directly from individual reads, and specific tools are not yet available. Another application is the detection of structural variants in cancer cells. 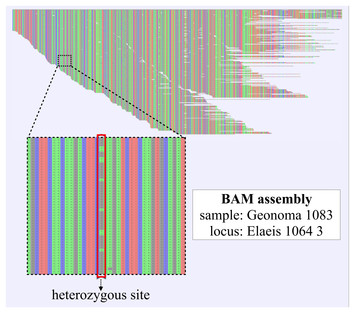
This is the case in directed evolution experiments, in which the genetic libraries typically originate from a single ancestral sequence (the wild-type) that has been submitted to limited randomization, for example, using error-prone PCR (epPCR). There is an increasing interest in using next-generation sequencing technologies for analyzing gene libraries that are highly diverse but have low variability (i.e., containing many different sequences differing from each other by only a few point mutations and for which a reference is available). Therefore, it is an attractive device for sequencing libraries of amplicons that are too long for other next-generation sequencing technologies. A minION device can read DNA strands of various lengths, from PCR products up to megabase genomic fragments, and current versions return at least 5 × 10 9 bases in one run. This approach offers portability and real-time sequencing, using simple experimental protocols, for a relatively low cost. It provides sequence base calls reconstructed from conductivity records during the translocation of a single DNA molecule through a protein pore. Nanopore is a powerful technology for high-throughput DNA sequencing, currently commercialized by Oxford Nanopore Technologies. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
